About GO-Module
About GO-Module
GO-Module is a web-accessible tool developed for
end-user biologists to greatly simplify the interpretation of prioritized Gene Ontology terms. GO-Module
radically reduces the complexity of raw Gene Ontology results creating compact biomodules in two distinct ways,
by 1) constructing biomodules from significant GO terms based on hierarchical knowledge, and 2) refining the GO
terms in each biomodule to contain only true positive results. Altogether, GO-Module outputs are an order
of magnitude smaller than the input set.
For details on why false positive signals can occur in functional enrichment studies of gene sets, see
Suppl. Fig. 8 and
Section B of Protocol
S1 in Citation 1 (Lee Yet al., 2010). In this head and neck cancer publication, our algorithm identified 0% to
59% of the total false positive enrichment results per gene set, with an average of 30%.
Tutorial
Figure1. Input fields
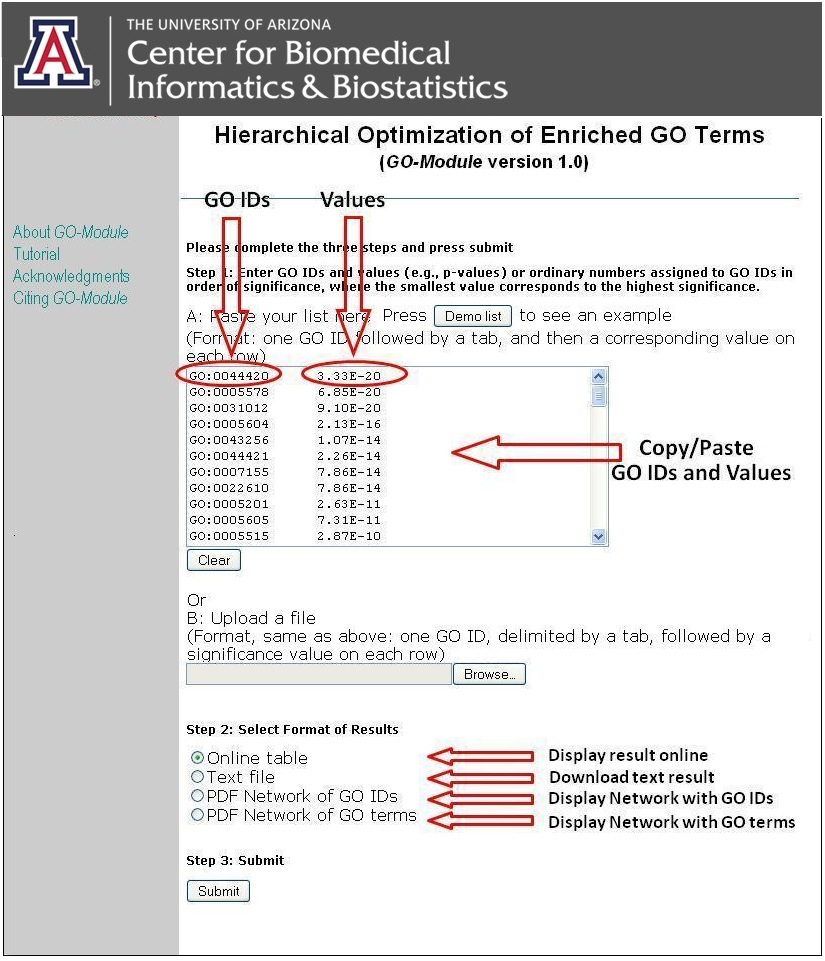
Figure2. Tabular output of GO-Module
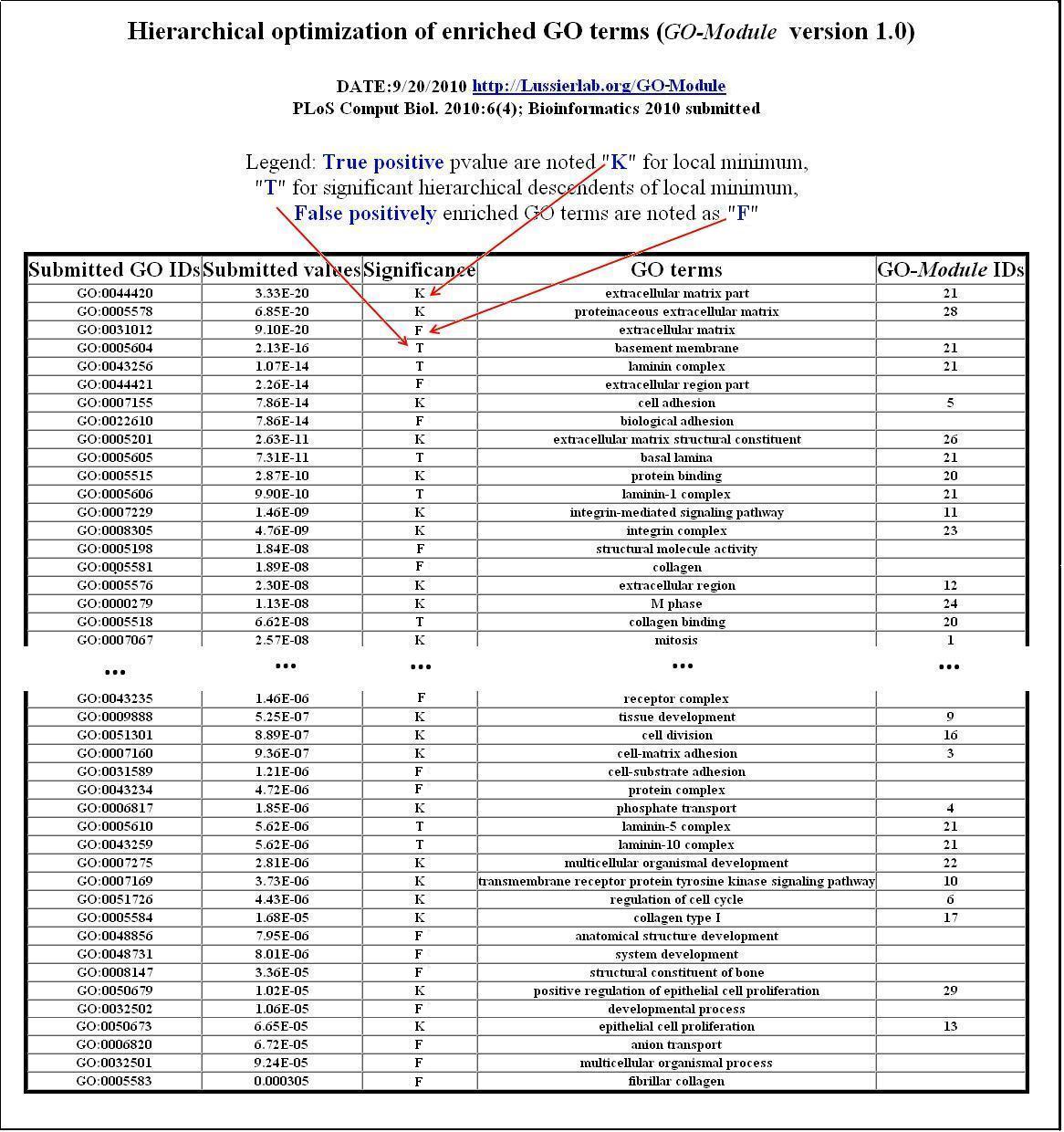
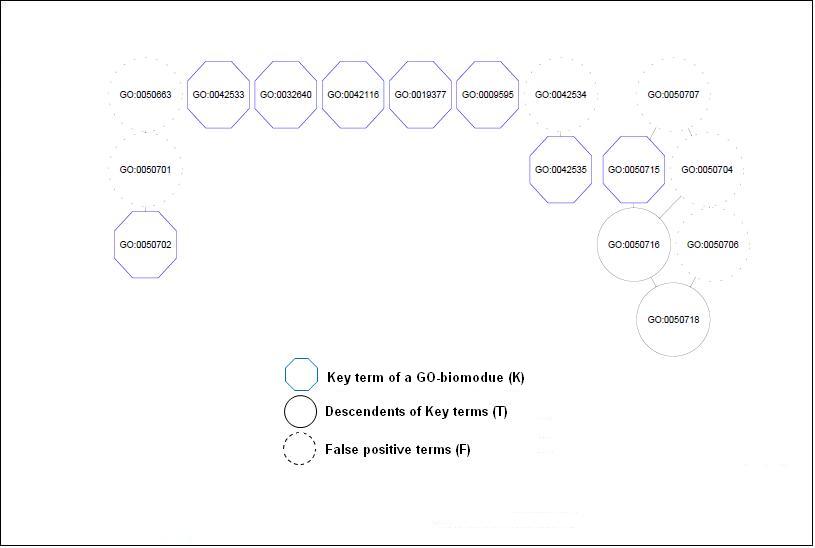
Figure3. Network output of GO-Module
Download Demo File
demofile.txt (Right-click the link and choose "Save Link As..." to save the fle to your computer)
Acknowledgments
This work was supported by NIH grants K22 LM008308, 1U54CA121852 and UL1 RR024999 and the Cancer Research
Foundation.
How to cite GO-Module
1. Yang X, Li J, Lee Y, Lussier YA*. GO-Module: functional synthesis and improved interpretation of Gene Ontology patterns. Bioinformatics 2011 Mar 17: 1444-14462 PMCID: PMC3087953.
Top